The Revolution in Single-Cell Image Analysis: From Algorithms to Agentic AI
Introduction: Beyond the Average Cell
In the era of precision medicine, the “average” cell—a concept central to traditional bulk RNA sequencing—is no longer sufficient to guide therapeutic decisions. Because tissues, particularly tumors, are highly heterogeneous, averaging the gene expression of thousands of cells masks critical information about rare subpopulations that drive disease progression and resistance.
Why does a therapy work for one patient but fail in another?
Why do genetically similar cells behave differently within the same tissue?
The answer lies at the single-cell level.
Single-cell image analysis has fundamentally transformed biological research—enabling scientists to observe cellular heterogeneity, spatial organization, and dynamic behavior with unprecedented resolution. However, as imaging technologies advance, they generate data of overwhelming complexity.
What began as rule-based image processing has evolved into intelligent, autonomous systems capable of reasoning across images, omics data, and biological knowledge.
We are witnessing a paradigm shift:
Traditional algorithms → Deep learning → Agentic AI
This blog traces the remarkable evolution of single-cell image analysis and explores how Agentic AI is redefining how we extract biological meaning from images and multi-omics data.
1. Understanding the True Complexity of Single-Cell Imaging
Single-cell image analysis is not merely about capturing images—it is about solving multi-scale biological puzzles.
a) Crowding: The Overlap Problem
In dense tissues such as solid tumors, cells are tightly packed and often intertwined. Traditional segmentation methods frequently suffer from segmentation leakage, where signals from one cell are misassigned to neighboring cells—leading to incorrect biological conclusions.
b) The 3D Reality
Cells exist in a three-dimensional microenvironment. Analyzing a 2D slice is like reading one page of a novel and guessing the entire plot.
Advanced imaging modalities such as:
- Light Sheet Fluorescence Microscopy (LSFM)
- Confocal and spinning disk microscopy
enable 3D visualization but produce terabytes of data per sample, far exceeding the capacity of traditional workflows.
c) Noise vs Signal: The Phototoxicity Paradox
To detect weak fluorescence signals, higher light intensity is required—but this causes:
- Photobleaching
- Phototoxicity
- Low-contrast, noisy images
Advanced denoising and restoration become mandatory before meaningful analysis can begin.
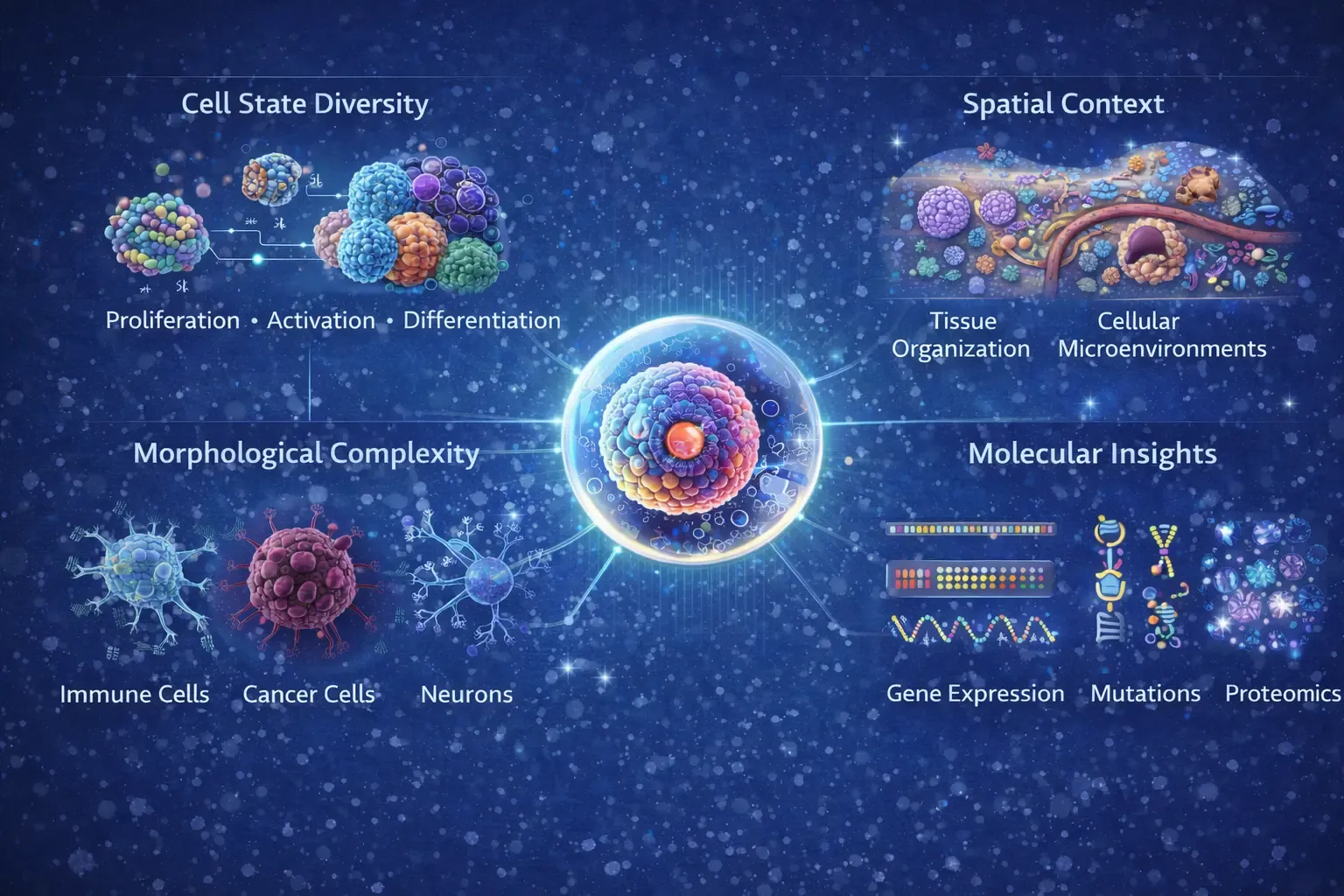
Figure 1: Figure depicts the intrinsic complexity of single-cell imaging. At the center, a single cell represents the fundamental analytical unit, surrounded by interconnected dimensions of cellular information. Cell state diversity captures dynamic biological processes such as proliferation, activation, and differentiation across heterogeneous cell populations. Morphological complexity highlights structural variability among immune cells, cancer cells, and neurons observed through high-resolution imaging. Spatial context illustrates tissue organization and cellular microenvironments that influence cell–cell interactions and functional states. Molecular insights integrate gene expression, mutational profiles, and proteomic signatures.
2. The Early Era: Handcrafted Algorithms & Classical Computer Vision
The earliest phase of single-cell image analysis relied on deterministic, rule-based pipelines.
Key Techniques
- Thresholding & filtering: Separating foreground from background
- Edge detection: Sobel, Canny, Laplacian operators
- Morphological operations: Erosion, dilation, opening, closing
- Watershed & region growing: Treating pixel intensity as topography
Toolboxes of the Era
- CellProfiler: Modular pipelines for high-content screening, requiring extensive parameter tuning
- Fiji/ImageJ: A flexible “Swiss Army knife” with thousands of plugins
- QuPath: Optimized for whole-slide imaging in digital pathology
- MATLAB toolboxes: Custom, dataset-specific solutions
Limitations
✔ Required extensive domain expertise
✔ Highly sensitive to staining and noise
✔ Rigid and poorly generalizable
These methods could detect cells, but they could not interpret biology.
3. Handcrafted Feature Engineering: A Bridge Between Vision and Biology
To improve biological relevance, researchers began extracting handcrafted features from segmented cells:
- Morphological: Area, perimeter, eccentricity
- Intensity-based: Mean, variance, texture
- Texture: Haralick features, Gabor filters
- Spatial: Nearest-neighbor distance, Voronoi tessellation
These features were fed into classical ML models:
- Support Vector Machines (SVMs)
- Random forests
- k-means clustering
While powerful, this approach still relied heavily on human intuition and struggled with biological variability.
4. The Deep Learning Revolution: The Great Enabler
The introduction of deep learning, particularly Convolutional Neural Networks (CNNs), marked a turning point.
Why Deep Learning Changed Everything
CNNs learn hierarchical features directly from raw pixels, eliminating manual feature engineering.
Key Breakthroughs
- U-Net: Pixel-level segmentation with encoder–decoder architecture
- Transfer learning: Leveraging ImageNet-trained models
- Multi-task learning: Segmentation, classification, and feature extraction in a single model
Advanced Architectures
- Mask R-CNN: Instance segmentation of overlapping cells
- GANs: Synthetic data generation and augmentation
- Vision Transformers (ViTs): Capturing long-range spatial dependencies via attention mechanism that focuses on biologically relevant regions
Key Deep Learning–Based Tools
Segmentation & Detection
-
Cellpose – Generalist deep learning model for cell and nucleus segmentation
-
StarDist – Star-convex polygon-based nuclear segmentation
-
DeepCell – CNN-based segmentation and tracking
-
U-Net variants – Backbone architecture across many tools
Interactive & Semi-Supervised Tools
-
Ilastik – ML-assisted pixel classification and segmentation
-
QuPath (DL extensions) – Digital pathology and tissue analysis
Impact
✔ Massive throughput
✔ Higher accuracy and reproducibility
✔ Discovery of subtle phenotypes
✔ Integration with multi-omics
But deep learning still lacked reasoning.
It learned patterns—but not biological intent.
5. The Rise of Multi-Modal Integration
Images provide structure—but omics provide molecular identity.
Why Integration Matters
True biological understanding requires:
- Morphology + gene expression
- Spatial context + molecular state
Emerging Capabilities
- Mapping scRNA-seq to spatial images
- Identifying spatial gene-expression gradients
- Linking cell identity to microenvironment
Representative Tools
- Tangram – mapping scRNA to spatial coordinates
- Cell2location – probabilistic cell type deconvolution
- SpaOTsc — optimal transport for spatial labeling
- Giotto / Squidpy — spatial patterns and statistics
Despite these advances, workflows remained manual, fragmented, and expert-driven.
6. The Next Frontier: Agentic AI
We are now entering the era of Agentic AI.
Unlike static pipelines, agents can observe, reason, and act autonomously.
What Makes AI “Agentic”?
- Goal-driven reasoning: Interprets natural-language research questions
- Autonomous planning: Designs multi-step analytical workflows
- Self-correction: Detects errors and adapts strategies
- Learning from feedback: Improves continuously with human input
Capabilities
- Switching segmentation models based on QC feedback
- Re-running denoising if image quality is poor
- Predicting molecular states from morphology alone
Real-World Highlight
TissueLab exemplifies a co-evolving agentic system:
- Scientists ask questions in natural language
- The agent generates explainable workflows
- Human corrections feed back into model improvement
Cellatria agentic AI tools in single-cell genomics analysis, along with how it works and why it matters for researchers:
- Automated Data Ingestion & Standardization
- Chatbot-Driven Interaction
- Agentic Workflow Execution
- Skill-Agnostic Access
7. Building Blocks: The Open-Source Agentic Stack
Agentic systems are powered by modular open-source frameworks:
- Coordinator / Planner: LangGraph-based workflow reasoning
- Tool Factory: Dynamically writes and tests new analysis code
- Critic Agent: Autonomous biological QC and validation
Together, they form a closed-loop intelligence system.
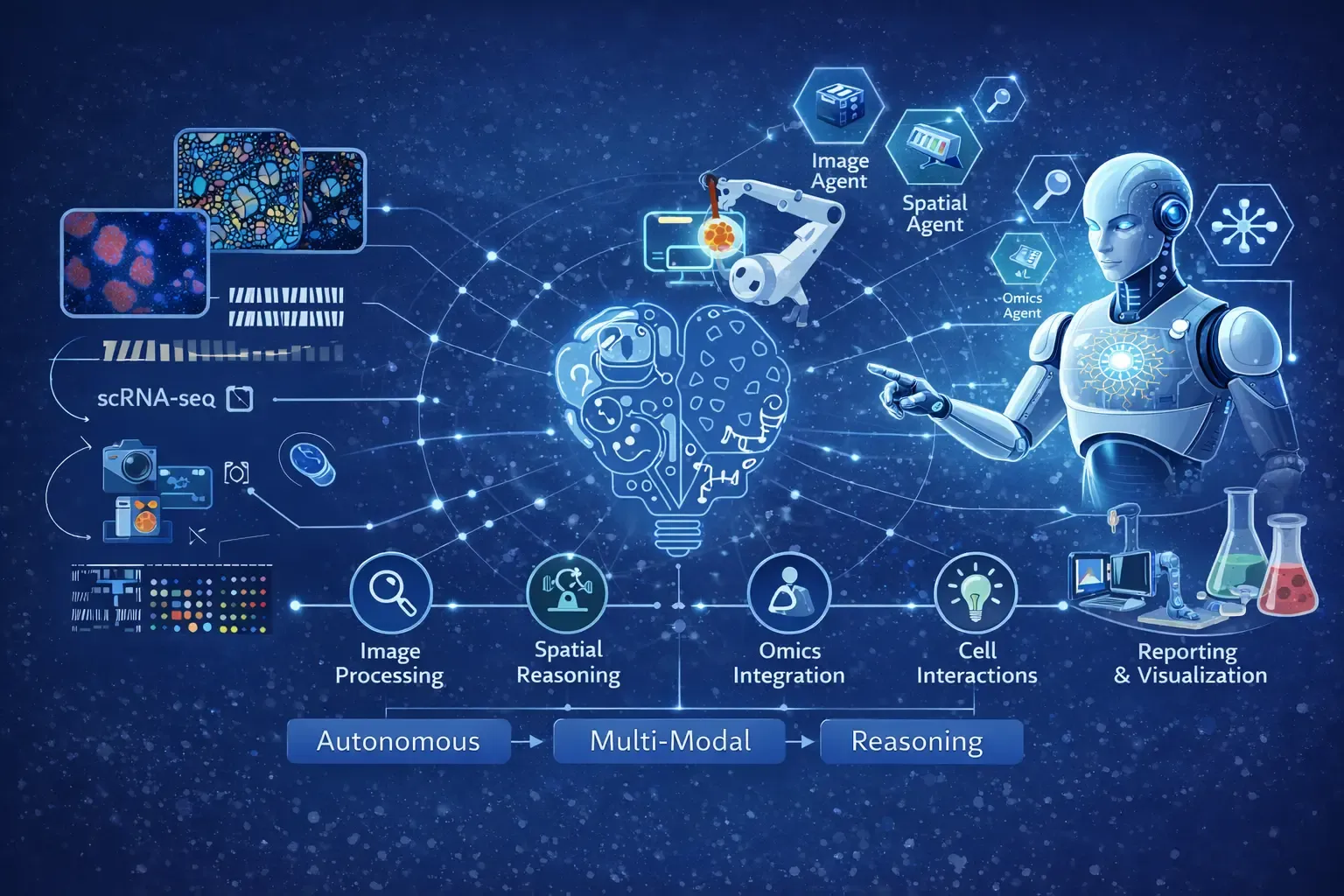
Figure 2: An Example of Agentic AI based Autonomous Reasoning Framework for Single-Cell Imaging. The figure illustrates how single-cell imaging data is interpreted through an autonomous, multimodal reasoning process. Individual cells serve as the core analytical units, capturing heterogeneity in cellular morphology, phenotypic states, and functional diversity. Spatial relationships represent tissue architecture and local microenvironments that govern cell–cell interactions. Molecular layers integrate gene expression, protein abundance, and signaling pathways to contextualize observed visual features. These interconnected modalities are processed through an autonomous reasoning flow, enabling agentic AI systems to dynamically integrate visual, spatial, and molecular information, uncover emergent cellular patterns, and generate biologically meaningful interpretations at single-cell resolution.
Conclusion: From Pixels to Biological Intelligence
The future of single-cell image analysis is not just about better images—it is about smarter interpretation.
We are moving toward:
- Agent-to-agent ecosystems
- Imaging agents collaborating with sequencing agents
- Fully integrated, explainable biological reports
This evolution mirrors the broader trajectory of AI in science:
From detection → to prediction → to reasoning → to autonomy
Agentic AI now bridges the gap between raw pixels and actionable biological insight, enabling discoveries at a scale previously unimaginable.
The revolution has begun—and the most exciting discoveries are yet to come.




